Enabling Python development on CDH clusters (for PySpark, for example) is now much easier thanks to new integration with Continuum Analytics’ Python platform (Anaconda).
Python has become an increasingly popular tool for data analysis, including data processing, feature engineering, machine learning, and visualization. Data scientists and data engineers enjoy Python’s rich numerical and analytical libraries—such as NumPy, pandas, and scikit-learn—and have long wanted to apply them to large datasets stored in Apache Hadoop clusters.
While Apache Spark, through PySpark, has made data in Hadoop clusters more accessible to Python users, actually using these libraries on a Hadoop cluster remains challenging. In particular, setting up a full-featured and modern Python environment on a cluster can be challenging, error-prone, and time-consuming.
For these reasons, Continuum Analytics and Cloudera have partnered to create an Anaconda parcel for CDH to enable simple distribution and installation of popular Python packages and their dependencies. Anaconda dramatically simplifies installation and management of popular Python packages and their dependencies, and this new parcel makes it easy for CDH users to deploy Anaconda across a Hadoop cluster for use in PySpark, Hadoop Streaming, and other contexts where Python is available and useful.
The newly available Anaconda parcel:
- Includes 300+ of the most popular Python packages
- Simplifies the installation of Anaconda across a CDH cluster
- Will be updated with each new Anaconda release.
In the remainder of this blog post, you’ll learn how to install and configure the Anaconda parcel, as well as explore an example of training a scikit-learn model on a single node and then using the model to make predictions on data in a cluster.
Installing the Anaconda Parcel
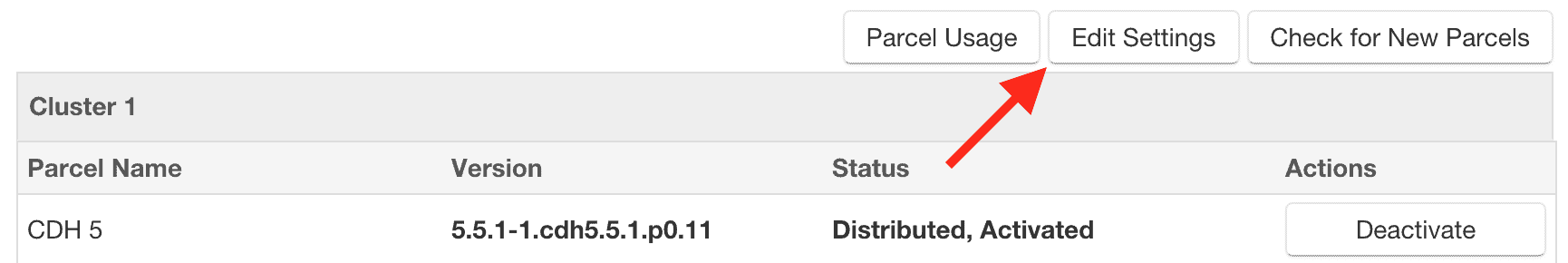
- From the Cloudera Manager Admin Console, click the “Parcels” indicator in the top navigation bar.
- Click the “Edit Settings” button on the top right of the Parcels page.
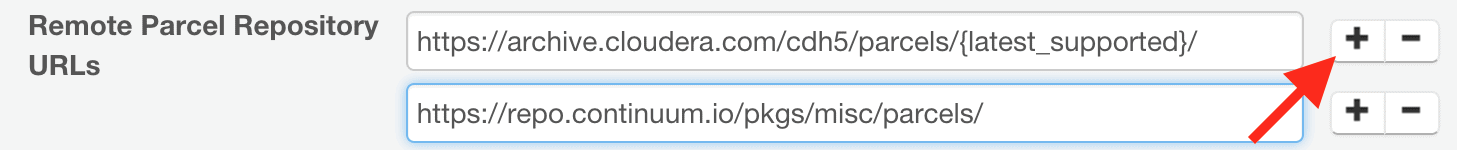
- Click the plus symbol in the “Remote Parcel Repository URLs” section, and add the following repository URL for the Anaconda parcel:
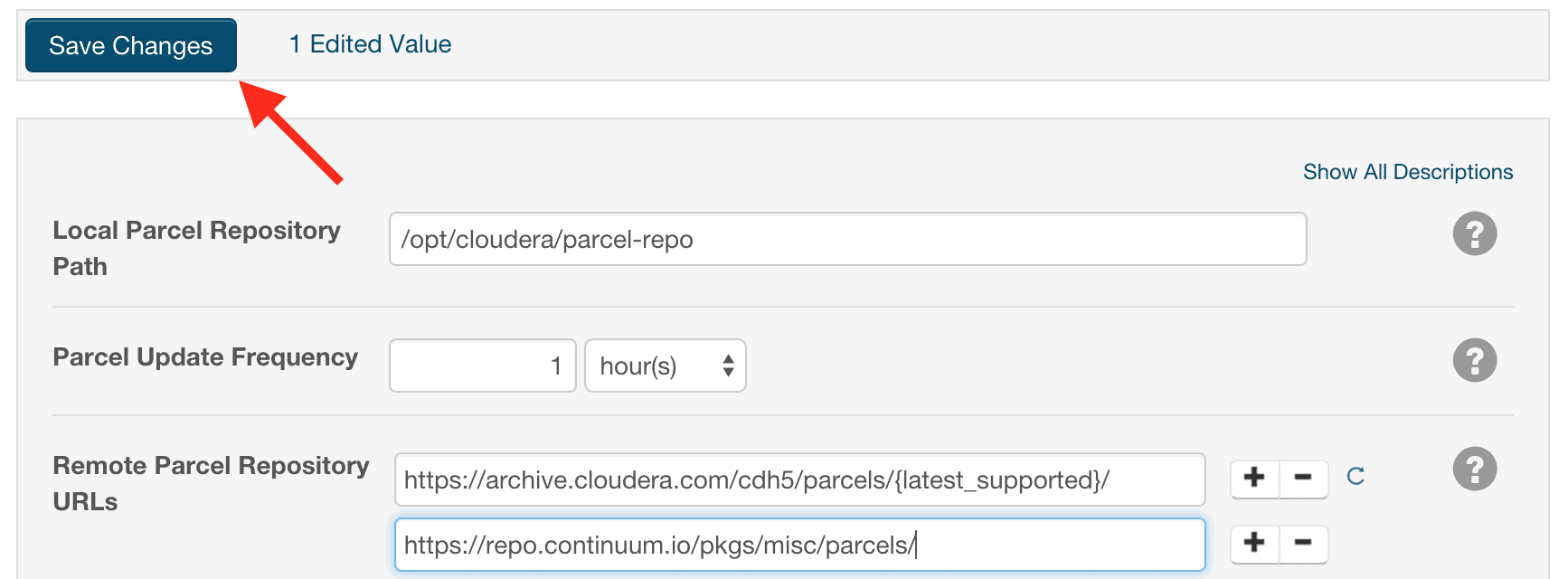
https://repo.continuum.io/pkgs/misc/parcels/ - Click the “Save Changes” button at the top of the page.
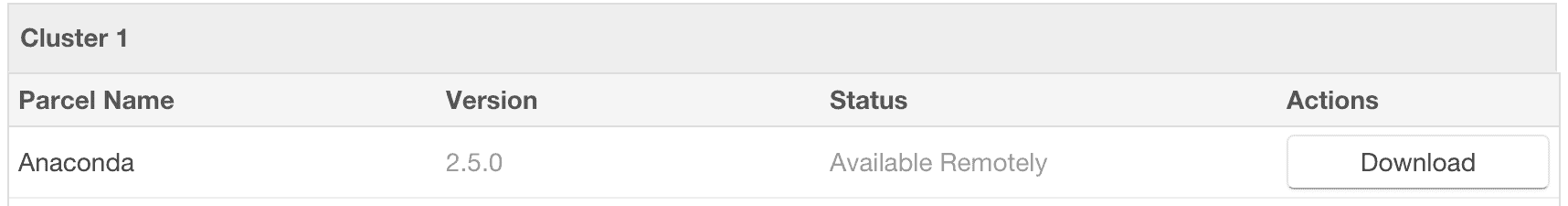
- Click the “Parcels” indicator in the top navigation bar to return to the list of available parcels, where you should see the latest version of the Anaconda parcel that is available.
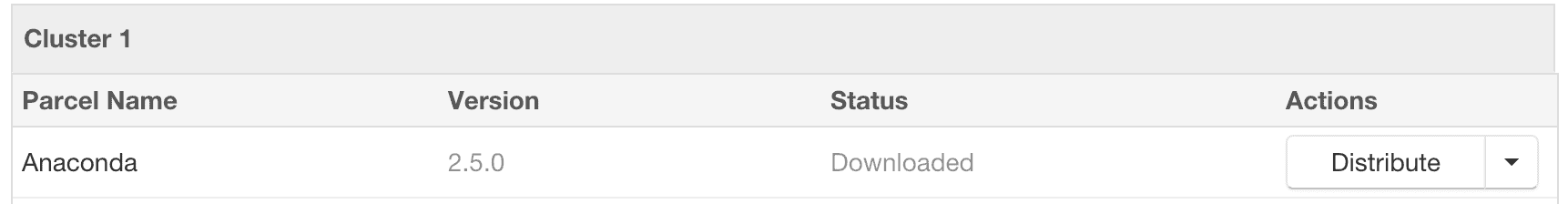
- Click the “Download” button to the right of the Anaconda parcel listing.
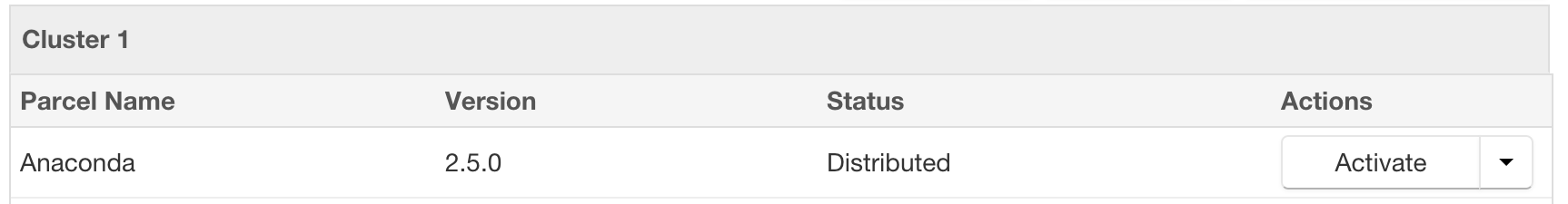
- After the parcel is downloaded, click the “Distribute” button to distribute the parcel to all of the cluster nodes.
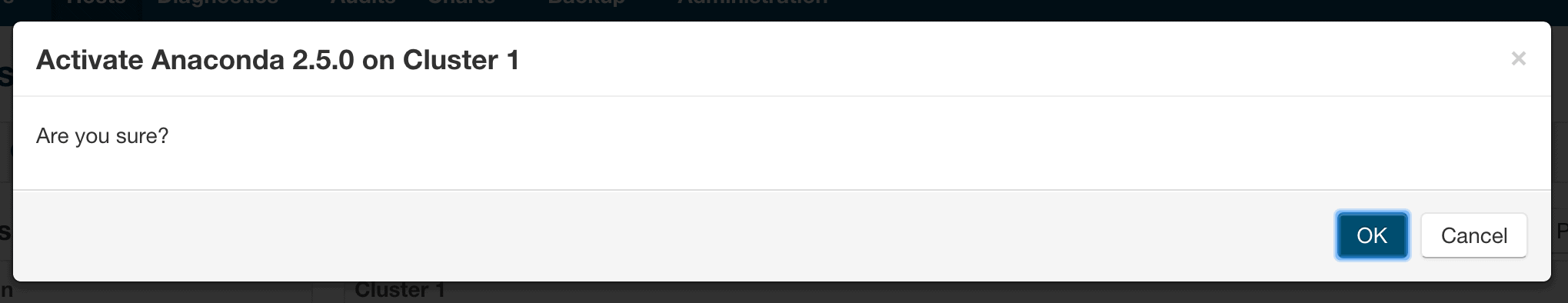
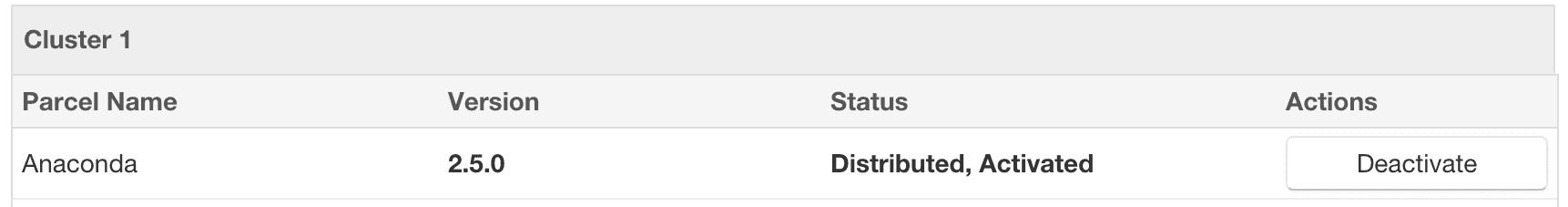
- After the parcel is distributed, click the “Activate” button to activate the parcel on all of the cluster nodes, which will prompt with a confirmation dialog.
- After the parcel is activated, Anaconda is now available on all of the cluster nodes.
These instructions are current as of the day of publication. Up-to-date instructions will be maintained in Anaconda’s documentation.
To make Spark aware that you want to use the installed parcels as the Python runtime environment on the cluster, you need to set the PYSPARK_PYTHON environment variable. Spark determines which Python interpreter to use by checking the value of the PYSPARK_PYTHON environment variable on the driver node. With the default configuration for Cloudera Manager and parcels, Anaconda will be installed to /opt/cloudera/parcels/Anaconda, but if the parcel directory for Cloudera Manager has been changed, you will need to change the below instructions to ${YOUR_PARCEL_DIR}/Anaconda/bin/python.
To specify which Python to use on a per-application basis, you can specify it on the same line as your spark-submit command. This would look like:
$ PYSPARK_PYTHON=/opt/cloudera/parcels/Anaconda/bin/python spark-submit pyspark_script.py
You can also use Anaconda by default in Spark applications while still allowing users to override the value if they wish. To do this, you will need follow the instructions for Advanced Configuration Snippets and add the following lines to Spark’s configuration:
if [ -z "${PYSPARK_PYTHON}" ]; then
export PYSPARK_PYTHON=/opt/cloudera/parcels/Anaconda/bin/python
fi
Now with Anaconda on your CDH cluster, there’s no need to manually install, manage, and provision Python packages on your Hadoop cluster.
Anaconda in Action
A commonly needed workflow for a Python using data scientist is to:
- Train a scikit-learn model on a single node.
- Save the results to disk.
- Apply the trained model using PySpark to generate predictions on a larger dataset.
Let’s take a classic machine-learning classification problem as an example of what having complex Python dependencies from Anaconda installed on CDH cluster allows you to do.
The MNIST dataset is a canonical machine-learning classification problem that involves recognizing handwritten digits, where each row of the dataset consists of a representation of one handwritten digit from 0 to 9. The training data you will use is the original MNIST dataset (60,000 rows). The prediction will be done with the MNIST8M dataset (8,000,000 rows). Both of these datasets are available from the libsvm datasets website. This dataset is used as a standard test for various machine-learning algorithms. More information, including benchmarks, can be found on the MNIST Dataset website.
To train the model on a single node, you will use scikit-learn and then save the model to a file with pickle:
with open('mnist.scale', 'r') as f:
train = f.read()
with open('mnist.scale.t', 'r') as f:
test = f.read()
import numpy as np
def parse(data):
lines = data.split('\n')
lines = filter(lambda x: x.strip() != '', lines)
nlines = len(lines)
X = np.zeros((nlines, 784) , dtype=float)
Y = np.zeros((nlines, ) , dtype=float)
for n, line in enumerate(lines):
line = line.strip()
if line != '':
parts = line.strip().split(' ')
for pair in parts[1:]:
pos, val = pair.strip().split(':')
pos, val = int(pos), float(val)
X[n, pos] = float(val)
Y[n] = parts[0]
return X, Y
X_train, Y_train = parse(train)
X_test, Y_test = parse(test)
from sklearn import svm, metrics
classifier = svm.SVC(gamma=0.001)
classifier.fit(X_train, Y_train)
predicted = classifier.predict(X_test)
print metrics.classification_report(Y_test, predicted)
import pickle
with open('classifier.pickle', 'w') as f:
pickle.dump(classifier, f)
With the classifier now trained, you can save it to disk and then copy it to HDFS.
Next, configure and create a SparkContext to run in yarn-client mode:
from pyspark import SparkConf
from pyspark import SparkContext
conf = SparkConf()
conf.setMaster('yarn-client')
conf.setAppName('sklearn-predict')
sc = SparkContext(conf=conf)
To load the MNIST8M data from HDFS into an RDD:
input_data = sc.textFile('hdfs:///tmp/mnist8m.scale')
Now let’s do some preprocessing on this dataset to convert the text to a NumPy array, which will serve as input for the scikit-learn classifier. You’ve installed Anaconda on every cluster node, so both NumPy and scikit-learn are available to the Spark worker processes.
def clean(line):
"""
Read the mnist8m file format and return a numpy array
"""
import numpy as np
X = np.zeros((1, 784) , dtype=float)
parts = line.strip().split(' ')
for pair in parts[1:]:
pos, val = pair.strip().split(':')
pos, val = int(pos), float(val)
if pos < pos:
X[0, pos] = float(val)
return X
inputs = input_data.map(clean)
To import the scikit-learn model and load the training data:
from sklearn.externals import joblib
classifier = joblib.load('classifier.pickle')
To apply the trained model to a data in a large file in HDFS, you need the trained model available in memory on the executors. To move the classifier from one node to all of the Spark workers, you can then use the SparkContext.broadcast function to:
broadcastVar = sc.broadcast(classifier)
This broadcast variable is then available in executors; thus you can use the variable in logic that needs to be executed on the cluster (inside of map or flatMap functions, for example). It is simple to apply the trained model and save the output to a file:
def apply_classifier(input_array):
label = broadcastVar.value.predict(input_array)
return label
predictions = inputs.map(apply_classifier)
predictions.saveAsTextFile('hdfs:///tmp/predictions')
To submit this code as a script, add the environment variable declaration at the beginning and then the usual spark-submit command:
PYSPARK_PYTHON=/opt/cloudera/parcels/Anaconda/bin/python spark-submit pyspark_job.py
Conclusion
Getting started with Anaconda on your CDH cluster is easy with the newly available parcel. Be sure to check out the Anaconda parcel documentation for more details; support is available through Continuum Analytics.
Juliet Hougland is a Data Scientist at Cloudera.
Kristopher Overholt is a Software Engineer at Continuum Analytics.
Daniel Rodriguez is a Data Scientist at Continuum Analytics.